Commit
·
1a1cab9
1
Parent(s):
35e3d0f
update readme
Browse files- README.md +95 -3
- assets/images/architecture.png +0 -0
README.md
CHANGED
|
@@ -10,10 +10,103 @@ license: other
|
|
| 10 |
|
| 11 |
# AIDO.StructureEncoder
|
| 12 |
|
| 13 |
-
AIDO.
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 14 |
|
| 15 |
## How to Use
|
| 16 |
-
Please see `experiments/AIDO.StructureTokenizer` in [Model Generator](https://github.com/genbio-ai/modelgenerator)
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 17 |
|
| 18 |
# Citation
|
| 19 |
Please cite AIDO.StructureTokenizer using the following BibTex code:
|
|
@@ -28,4 +121,3 @@ Please cite AIDO.StructureTokenizer using the following BibTex code:
|
|
| 28 |
booktitle={NeurIPS 2024 Workshop on AI for New Drug Modalities},
|
| 29 |
}
|
| 30 |
```
|
| 31 |
-
- Docs: [More Information Needed]
|
|
|
|
| 10 |
|
| 11 |
# AIDO.StructureEncoder
|
| 12 |
|
| 13 |
+
AIDO.StructureEncoder is the encoder-only component of [AIDO.StructureTokenizer](https://huggingface.co/genbio-ai/AIDO.StructureTokenizer) for tokenization of protein structures.
|
| 14 |
+
|
| 15 |
+
## Model Description
|
| 16 |
+
|
| 17 |
+

|
| 18 |
+
|
| 19 |
+
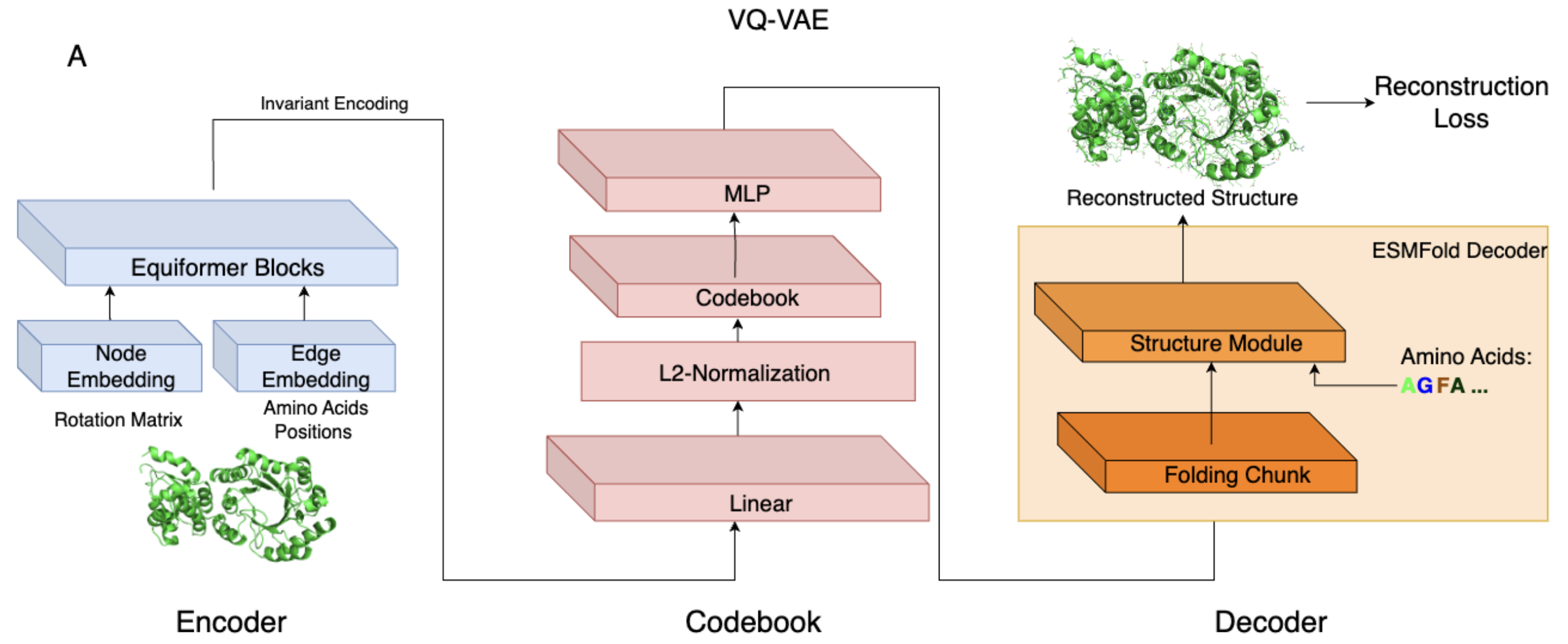
**AIDO.StructureTokenizer** is built on a Vector Quantized Variational Autoencoder (VQ-VAE) architecture with the following components:
|
| 20 |
+
- Equivariant Encoder (6M): Encodes backbone structures into a latent space that maintains rotational and translational symmetries using the Equiformer architecture.
|
| 21 |
+
- Discrete Codebook: Maps continuous latent vectors into 512 discrete structural tokens.
|
| 22 |
+
- Invariant Decoder (300M): Reconstructs full 3D structures, including side chains, from the structural tokens using an architecture adapted from ESMFold.
|
| 23 |
+
|
| 24 |
+
This model strikes a balance between reconstruction fidelity and structural locality, optimizing its suitability for downstream tasks such as structure prediction, homology detection, and multimodal protein modeling.
|
| 25 |
+
|
| 26 |
+
### Key Features
|
| 27 |
+
|
| 28 |
+
- Encoding Structures into Tokens (See [below](#how-to-use))
|
| 29 |
+
- Decoding Tokens into Structures (See [genbio-ai/AIDO.StructureDecoder](https://huggingface.co/genbio-ai/AIDO.StructureDecoder))
|
| 30 |
+
- Reconstructing Structures (See [genbio-ai/AIDO.StructureTokenizer](https://huggingface.co/genbio-ai/AIDO.StructureTokenizer))
|
| 31 |
+
- Structure Prediction (See [this section](https://huggingface.co/genbio-ai/AIDO.Protein2StructureToken-16B/blob/main/README.md#structure-prediction) in genbio-ai/AIDO.Protein2StructureToken-16B)
|
| 32 |
+
|
| 33 |
|
| 34 |
## How to Use
|
| 35 |
+
Please see `experiments/AIDO.StructureTokenizer` in [Model Generator](https://github.com/genbio-ai/modelgenerator) for more details.
|
| 36 |
+
|
| 37 |
+
### Setup
|
| 38 |
+
Install [Model Generator](https://github.com/genbio-ai/modelgenerator)
|
| 39 |
+
|
| 40 |
+
#### Data preparation
|
| 41 |
+
|
| 42 |
+
To reproduce the reconstruction results in the paper, we provide a preprocessed CASP15 dataset at [genbio-ai/sample-structure-dataset](https://huggingface.co/datasets/genbio-ai/sample-structure-dataset). It could be downloaded via
|
| 43 |
+
```bash
|
| 44 |
+
huggingface-cli download genbio-ai/sample-structure-dataset --repo-type dataset --local-dir ./data/protstruct_sample_data/
|
| 45 |
+
```
|
| 46 |
+
|
| 47 |
+
This dataset is based on the CASP15 dataset, which can be referenced at:
|
| 48 |
+
- [CASP15 Prediction Center](https://predictioncenter.org/casp15/)
|
| 49 |
+
- [Bhattacharya-Lab/CASP15](https://github.com/Bhattacharya-Lab/CASP15)
|
| 50 |
+
|
| 51 |
+
The downloaded directory includes:
|
| 52 |
+
- A `registries` folder containing a CSV file with metadata such as filenames and PDB IDs.
|
| 53 |
+
- A `CASP15_merged` folder containing PDB files, where domains are merged in the same way as described in [Bhattacharya-Lab/CASP15](https://github.com/Bhattacharya-Lab/CASP15).
|
| 54 |
+
|
| 55 |
+
To use customized data, you can prepare a dataset with the following structure:
|
| 56 |
+
- A folder containing PDB files (supported formats: `cif.gz`, `cif`, `ent.gz`, `pdb`).
|
| 57 |
+
|
| 58 |
+
Then, you need to prepare a registry file in CSV format using the following command:
|
| 59 |
+
``` bash
|
| 60 |
+
python experiments/AIDO.StructureTokenizer/register_dataset.py \
|
| 61 |
+
--folder_path /path/to/folder_path \
|
| 62 |
+
--format cif.gz \
|
| 63 |
+
--output_file /path/to/output_file.csv
|
| 64 |
+
```
|
| 65 |
+
|
| 66 |
+
You need to replace the `folder_path` and the `registry_path` in the following steps accordingly.
|
| 67 |
+
|
| 68 |
+
#### Running Encoding Task
|
| 69 |
+
|
| 70 |
+
If you use the sample dataset, you can run the encoding task using the following command:
|
| 71 |
+
```bash
|
| 72 |
+
CUDA_VISIBLE_DEVICES=0 mgen predict --config experiments/AIDO.StructureTokenizer/encode.yaml
|
| 73 |
+
```
|
| 74 |
+
|
| 75 |
+
If you use your own dataset, you need to update the `folder_path` and the `registry_path` in the `encode.yaml` configuration file to point to your dataset folder and registry file. Alternatively, you can override these parameters when running the command:
|
| 76 |
+
```bash
|
| 77 |
+
CUDA_VISIBLE_DEVICES=0 mgen predict --config experiments/AIDO.StructureTokenizer/encode.yaml \
|
| 78 |
+
--data.init_args.config.proteins_datasets_configs.name="your_dataset_name" \
|
| 79 |
+
--data.init_args.config.proteins_datasets_configs.registry_path="your_dataset_folder_path" \
|
| 80 |
+
--data.init_args.config.proteins_datasets_configs.folder_path="your_dataset_registry_path" \
|
| 81 |
+
--trainer.callbacks.dict_kwargs.output_dir="your_output_dir"
|
| 82 |
+
```
|
| 83 |
+
|
| 84 |
+
**Input:**
|
| 85 |
+
- The PDB files in the dataset folder.
|
| 86 |
+
- The registry file in CSV format indicating the metadata of the dataset.
|
| 87 |
+
|
| 88 |
+
**Output:**
|
| 89 |
+
- The encoded tokens will be saved in the output directory specified in the configuration file. By default it is saved in `logs/protstruct_encode/`.
|
| 90 |
+
- The encoded tokens are saved as a `.pt` file, which can be loaded using PyTorch. Inside the file, it's a dictionary that maps the name of the protein to the encoded tokens (`struct_tokens`) and other auxiliary information (`aatype`, `residue_index`) for reconstruction.
|
| 91 |
+
The structure of the dictionary is as follows:
|
| 92 |
+
```python
|
| 93 |
+
{
|
| 94 |
+
'T1137s5_nan': { # the nan here is the chain id and CASP15 doesn't have chain id
|
| 95 |
+
'struct_tokens': tensor([449, 313, 207, 129, ...]),
|
| 96 |
+
'aatype': tensor([ 4, 7, 5, 17, ...]),
|
| 97 |
+
'residue_index': tensor([ 33, 34, 35, 36, ...]),
|
| 98 |
+
},
|
| 99 |
+
...
|
| 100 |
+
}
|
| 101 |
+
```
|
| 102 |
+
- A codebook file (`codebook.pt`) that contains the embedding of each token. The shape is `(num_tokens, embedding_dim)`.
|
| 103 |
+
|
| 104 |
+
**Notes:**
|
| 105 |
+
- Currently, this function only supports single GPU inference due to the file saving mechanism. We plan to support multi-GPU inference in the future.
|
| 106 |
+
- The auxiliary information (`aatype` and `residue_index`) can be substituted with placeholder values if not required.
|
| 107 |
+
- `aatype`: This parameter is used to reconstruct the protein sidechains. If sidechain reconstruction is not needed, you can assign dummy values (e.g., all zeros).
|
| 108 |
+
- `residue_index`: This parameter helps the model identify gaps in residue numbering, which can influence structural predictions. If gaps are present, the model may introduce holes in the structure. For structures without gaps, you can use a continuous sequence of integers (e.g., 0 to n-1).
|
| 109 |
+
- You may need to adjust the `max_nb_res` parameter in the configuration file based on the maximum number of residues in your dataset. For those proteins with more residues than `max_nb_res`, the model will truncate the residues. The default value is set to 1024.
|
| 110 |
|
| 111 |
# Citation
|
| 112 |
Please cite AIDO.StructureTokenizer using the following BibTex code:
|
|
|
|
| 121 |
booktitle={NeurIPS 2024 Workshop on AI for New Drug Modalities},
|
| 122 |
}
|
| 123 |
```
|
|
|
assets/images/architecture.png
ADDED
|