Advancing Open-source Large Language Models in Medical Domain
Online Demo
|
 GitHub
|
GitHub
|
 Paper
|
Paper
|
 Discord
Discord
Introducing OpenBioLLM-70B: A State-of-the-Art Open Source Biomedical Large Language Model
OpenBioLLM-70B is an advanced open source language model designed specifically for the biomedical domain. Developed by Saama AI Labs, this model leverages cutting-edge techniques to achieve state-of-the-art performance on a wide range of biomedical tasks.
🏥 Biomedical Specialization: OpenBioLLM-70B is tailored for the unique language and knowledge requirements of the medical and life sciences fields. It was fine-tuned on a vast corpus of high-quality biomedical data, enabling it to understand and generate text with domain-specific accuracy and fluency.
🎓 Superior Performance: With 70 billion parameters, OpenBioLLM-70B outperforms other open source biomedical language models of similar scale. It has also demonstrated better results compared to larger proprietary & open-source models like GPT-4, Gemini, Meditron-70B, Med-PaLM-1 & Med-PaLM-2 on biomedical benchmarks.
🧠 Advanced Training Techniques: OpenBioLLM-70B builds upon the powerful foundations of the Meta-Llama-3-70B-Instruct and Meta-Llama-3-70B-Instruct models. It incorporates the DPO dataset and fine-tuning recipe along with a custom diverse medical instruction dataset. Key components of the training pipeline include:

- Policy Optimization: Direct Preference Optimization: Your Language Model is Secretly a Reward Model (DPO)
- Fine-tuning dataset: Custom Medical Instruct dataset (We plan to release a sample training dataset in our upcoming paper; please stay updated)
This combination of cutting-edge techniques enables OpenBioLLM-70B to align with key capabilities and preferences for biomedical applications.
⚙️ Release Details:
- Model Size: 70 billion parameters
- Quantization: Optimized quantized versions available Here
- Language(s) (NLP): en
- Developed By: Ankit Pal (Aaditya Ura) from Saama AI Labs
- License: Meta-Llama License
- Fine-tuned from models: Meta-Llama-3-70B-Instruct
- Resources for more information:
- Paper: Coming soon
The model can be fine-tuned for more specialized tasks and datasets as needed.
OpenBioLLM-70B represents an important step forward in democratizing advanced language AI for the biomedical community. By leveraging state-of-the-art architectures and training techniques from leading open source efforts like Llama-3, we have created a powerful tool to accelerate innovation and discovery in healthcare and the life sciences.
We are excited to share OpenBioLLM-70B with researchers and developers around the world.
Community & Resources
🔥 Your Daily Dose of Medical AI Breakthroughs 🚀
We turn hours of the latest research papers into minutes. Get daily tweets and news on the latest medical AI breakthroughs, dataset releases, and benchmark results – all carefully curated to save you time while keeping you informed.

Daily updates on Medical LLMs, datasets & benchmarks |

Daily news on Medical LLMs, datasets & benchmarks |

Video & audio summaries of latest research |

Connect with researchers & discuss latest developments |
Use with transformers
Important: Please use the exact chat template provided by Llama-3 instruct version. Otherwise there will be a degradation in the performance. The model output can be verbose in rare cases. Please consider setting temperature = 0 to make this happen less.
See the snippet below for usage with Transformers:
import transformers
import torch
model_id = "aaditya/OpenBioLLM-Llama3-70B"
pipeline = transformers.pipeline(
"text-generation",
model=model_id,
model_kwargs={"torch_dtype": torch.bfloat16},
device="auto",
)
messages = [
{"role": "system", "content": "You are an expert and experienced from the healthcare and biomedical domain with extensive medical knowledge and practical experience. Your name is OpenBioLLM, and you were developed by Saama AI Labs. who's willing to help answer the user's query with explanation. In your explanation, leverage your deep medical expertise such as relevant anatomical structures, physiological processes, diagnostic criteria, treatment guidelines, or other pertinent medical concepts. Use precise medical terminology while still aiming to make the explanation clear and accessible to a general audience."},
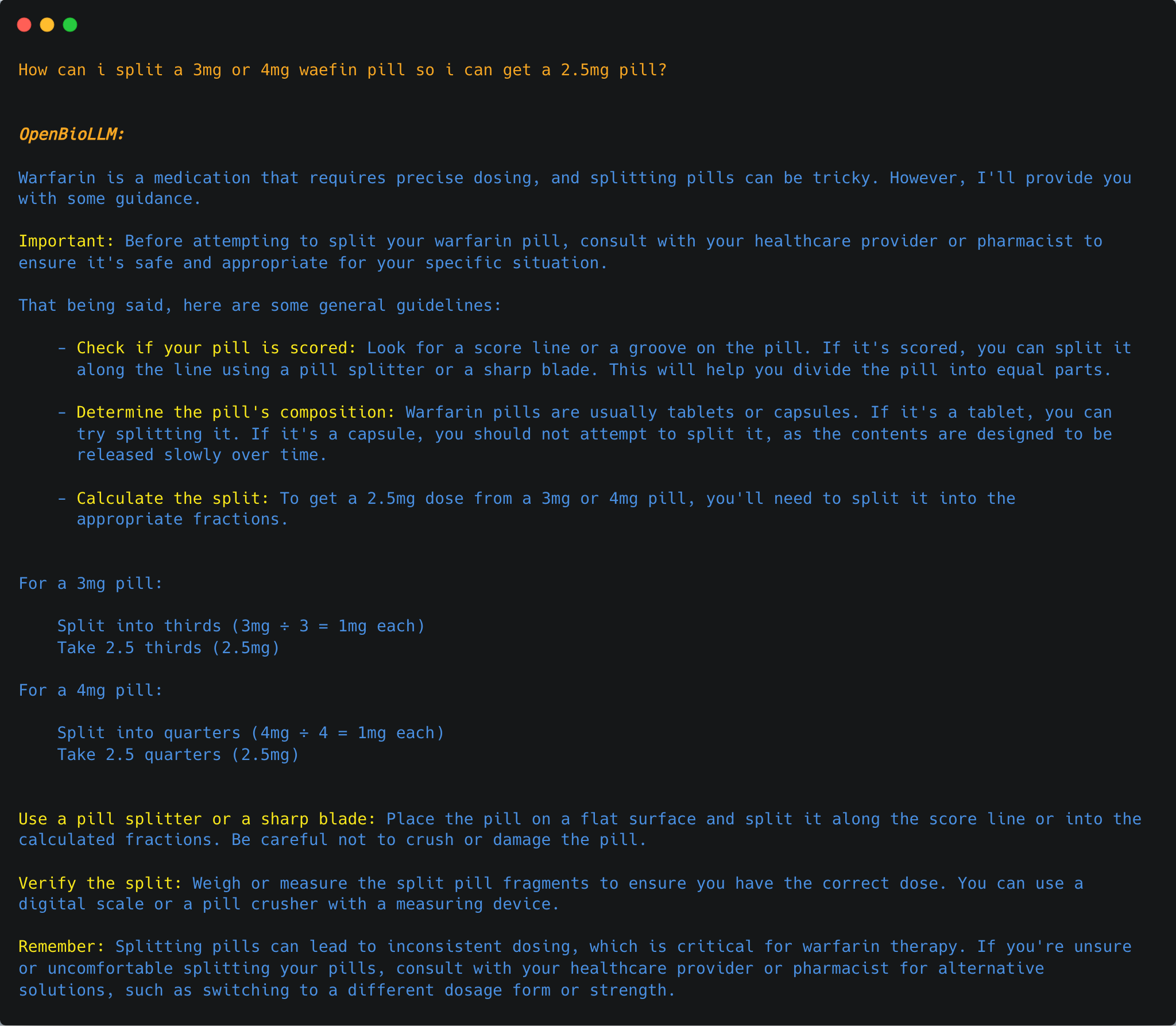
{"role": "user", "content": "How can i split a 3mg or 4mg waefin pill so i can get a 2.5mg pill?"},
]
prompt = pipeline.tokenizer.apply_chat_template(
messages,
tokenize=False,
add_generation_prompt=True
)
terminators = [
pipeline.tokenizer.eos_token_id,
pipeline.tokenizer.convert_tokens_to_ids("<|eot_id|>")
]
outputs = pipeline(
prompt,
max_new_tokens=256,
eos_token_id=terminators,
do_sample=True,
temperature=0.0,
top_p=0.9,
)
print(outputs[0]["generated_text"][len(prompt):])
Training procedure
Training hyperparameters
Click to see details
- learning_rate: 0.0002
- lr_scheduler: cosine
- train_batch_size: 12
- eval_batch_size: 8
- GPU: H100 80GB SXM5
- num_devices: 8
- optimizer: adamw_bnb_8bit
- lr_scheduler_warmup_steps: 100
- num_epochs: 4
Peft hyperparameters
Click to see details
- adapter: qlora
- lora_r: 128
- lora_alpha: 256
- lora_dropout: 0.05
- lora_target_linear: true
-lora_target_modules:
- q_proj
- v_proj
- k_proj
- o_proj
- gate_proj
- down_proj
- up_proj
Training results
Framework versions
- Transformers 4.39.3
- Pytorch 2.1.2+cu121
- Datasets 2.18.0
- Tokenizers 0.15.1
- Axolotl
- Lm harness for evaluation
Benchmark Results
🔥 OpenBioLLM-70B demonstrates superior performance compared to larger models, such as GPT-4, Gemini, Meditron-70B, Med-PaLM-1 & Med-PaLM-2 across 9 diverse biomedical datasets, achieving state-of-the-art results with an average score of 86.06%, despite having a significantly smaller parameter count. The model's strong performance in domain-specific tasks, such as Clinical KG, Medical Genetics, and PubMedQA, highlights its ability to effectively capture and apply biomedical knowledge.
🚨 The GPT-4, Med-PaLM-1, and Med-PaLM-2 results are taken from their official papers. Since Med-PaLM doesn't provide zero-shot accuracy, we are using 5-shot accuracy from their paper for comparison. All results presented are in the zero-shot setting, except for Med-PaLM-2 and Med-PaLM-1, which use 5-shot accuracy.
| Clinical KG | Medical Genetics | Anatomy | Pro Medicine | College Biology | College Medicine | MedQA 4 opts | PubMedQA | MedMCQA | Avg | |
|---|---|---|---|---|---|---|---|---|---|---|
| OpenBioLLM-70B | 92.93 | 93.197 | 83.904 | 93.75 | 93.827 | 85.749 | 78.162 | 78.97 | 74.014 | 86.05588 |
| Med-PaLM-2 (5-shot) | 88.3 | 90 | 77.8 | 95.2 | 94.4 | 80.9 | 79.7 | 79.2 | 71.3 | 84.08 |
| GPT-4 | 86.04 | 91 | 80 | 93.01 | 95.14 | 76.88 | 78.87 | 75.2 | 69.52 | 82.85 |
| Med-PaLM-1 (Flan-PaLM, 5-shot) | 80.4 | 75 | 63.7 | 83.8 | 88.9 | 76.3 | 67.6 | 79 | 57.6 | 74.7 |
| OpenBioLLM-8B | 76.101 | 86.1 | 69.829 | 78.21 | 84.213 | 68.042 | 58.993 | 74.12 | 56.913 | 72.502 |
| Gemini-1.0 | 76.7 | 75.8 | 66.7 | 77.7 | 88 | 69.2 | 58 | 70.7 | 54.3 | 70.79 |
| GPT-3.5 Turbo 1106 | 74.71 | 74 | 72.79 | 72.79 | 72.91 | 64.73 | 57.71 | 72.66 | 53.79 | 66 |
| Meditron-70B | 66.79 | 69 | 53.33 | 71.69 | 76.38 | 63 | 57.1 | 76.6 | 46.85 | 64.52 |
| gemma-7b | 69.81 | 70 | 59.26 | 66.18 | 79.86 | 60.12 | 47.21 | 76.2 | 48.96 | 64.18 |
| Mistral-7B-v0.1 | 68.68 | 71 | 55.56 | 68.38 | 68.06 | 59.54 | 50.82 | 75.4 | 48.2 | 62.85 |
| Apollo-7B | 62.26 | 72 | 61.48 | 69.12 | 70.83 | 55.49 | 55.22 | 39.8 | 53.77 | 60 |
| MedAlpaca-7b | 57.36 | 69 | 57.04 | 67.28 | 65.28 | 54.34 | 41.71 | 72.8 | 37.51 | 58.03 |
| BioMistral-7B | 59.9 | 64 | 56.5 | 60.4 | 59 | 54.7 | 50.6 | 77.5 | 48.1 | 57.3 |
| AlpaCare-llama2-7b | 49.81 | 49 | 45.92 | 33.82 | 50 | 43.35 | 29.77 | 72.2 | 34.42 | 45.36 |
| ClinicalGPT | 30.56 | 27 | 30.37 | 19.48 | 25 | 24.27 | 26.08 | 63.8 | 28.18 | 30.52 |
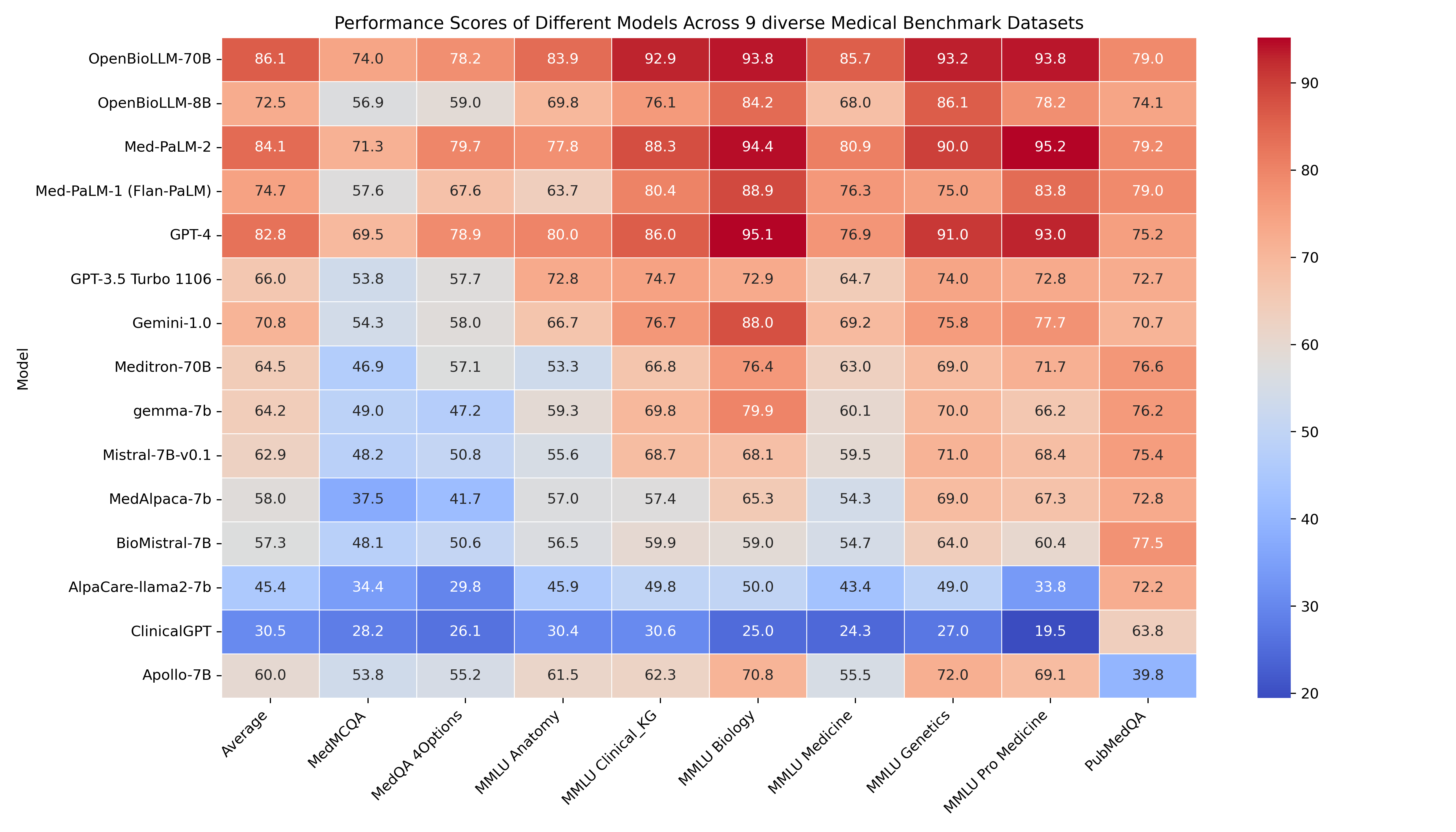
Detailed Medical Subjectwise accuracy
Use Cases & Examples
🚨 **Below results are from the quantized version of OpenBioLLM-70B
Summarize Clinical Notes
OpenBioLLM-70B can efficiently analyze and summarize complex clinical notes, EHR data, and discharge summaries, extracting key information and generating concise, structured summaries
Answer Medical Questions
OpenBioLLM-70B can provide answers to a wide range of medical questions.
Clinical Entity Recognition
OpenBioLLM-70B can perform advanced clinical entity recognition by identifying and extracting key medical concepts, such as diseases, symptoms, medications, procedures, and anatomical structures, from unstructured clinical text. By leveraging its deep understanding of medical terminology and context, the model can accurately annotate and categorize clinical entities, enabling more efficient information retrieval, data analysis, and knowledge discovery from electronic health records, research articles, and other biomedical text sources. This capability can support various downstream applications, such as clinical decision support, pharmacovigilance, and medical research.
Biomarkers Extraction
Classification
OpenBioLLM-70B can perform various biomedical classification tasks, such as disease prediction, sentiment analysis, medical document categorization
De-Identification
OpenBioLLM-70B can detect and remove personally identifiable information (PII) from medical records, ensuring patient privacy and compliance with data protection regulations like HIPAA.
Advisory Notice!
While OpenBioLLM-70B leverages high-quality data sources, its outputs may still contain inaccuracies, biases, or misalignments that could pose risks if relied upon for medical decision-making without further testing and refinement. The model's performance has not yet been rigorously evaluated in randomized controlled trials or real-world healthcare environments.
Therefore, we strongly advise against using OpenBioLLM-70B for any direct patient care, clinical decision support, or other professional medical purposes at this time. Its use should be limited to research, development, and exploratory applications by qualified individuals who understand its limitations. OpenBioLLM-70B is intended solely as a research tool to assist healthcare professionals and should never be considered a replacement for the professional judgment and expertise of a qualified medical doctor.
Appropriately adapting and validating OpenBioLLM-70B for specific medical use cases would require significant additional work, potentially including:
- Thorough testing and evaluation in relevant clinical scenarios
- Alignment with evidence-based guidelines and best practices
- Mitigation of potential biases and failure modes
- Integration with human oversight and interpretation
- Compliance with regulatory and ethical standards
Always consult a qualified healthcare provider for personal medical needs.
Citation
If you find OpenBioLLM-70B & 8B useful in your work, please cite the model as follows:
@misc{OpenBioLLMs,
author = {Ankit Pal, Malaikannan Sankarasubbu},
title = {OpenBioLLMs: Advancing Open-Source Large Language Models for Healthcare and Life Sciences},
year = {2024},
publisher = {Hugging Face},
journal = {Hugging Face repository},
howpublished = {\url{https://huggingface.co/aaditya/OpenBioLLM-Llama3-70B}}
}
The accompanying paper is currently in progress and will be released soon.
💌 Contact
We look forward to hearing you and collaborating on this exciting project!
Contributors:
- Ankit Pal (Aaditya Ura) [aadityaura at gmail dot com]
- Saama AI Labs
- Note: I am looking for a funded PhD opportunity, especially if it fits my Responsible Generative AI, Multimodal LLMs, Geometric Deep Learning, and Healthcare AI skillset.
References
We thank the Meta Team for their amazing models!
Result sources
- [1] GPT-4 [Capabilities of GPT-4 on Medical Challenge Problems] (https://arxiv.org/abs/2303.13375)
- [2] Med-PaLM-1 Large Language Models Encode Clinical Knowledge
- [3] Med-PaLM-2 Towards Expert-Level Medical Question Answering with Large Language Models
- [4] Gemini-1.0 Gemini Goes to Med School
- Downloads last month
- 23,319
Model tree for aaditya/Llama3-OpenBioLLM-70B
Base model
meta-llama/Meta-Llama-3-70B